
Nature Machine Intelligence Article
Optimizing molecular resource utilization to accelerate the molecular discovery process requires collaborative efforts among research institutions and organizations. However, given that both successful and failed molecules generated by each institution hold high research value, these discoveries are typically kept highly confidential before formal publication or commercialization. This confidentiality requirement poses significant challenges for most existing methods to collaboratively process heterogeneously distributed molecular data under strict privacy constraints. The University of Science and Technology of China (USTC) team proposed FedLG (Federated Learning with Lanczos Graph), a federated graph learning method that utilizes the Lanczos algorithm to facilitate multi-party collaborative model training, achieving reliable predictive performance under strict privacy protection conditions.
I. Research Background: The Paradox Between Molecular Data Privacy and Collaboration Needs
Molecular discovery is considered a key component driving the development of related fields such as materials design and drug discovery. With the advancement of artificial intelligence and experimental technologies, researchers now have access to more unique molecular structures. These molecules are typically distributed across different research institutions or organizations, presenting heterogeneous data distributions while being subject to strict privacy and confidentiality constraints. Open-access molecular databases represent only a small fraction of available molecular knowledge, as they cannot access high-value and confidential molecular data within private research institutions.
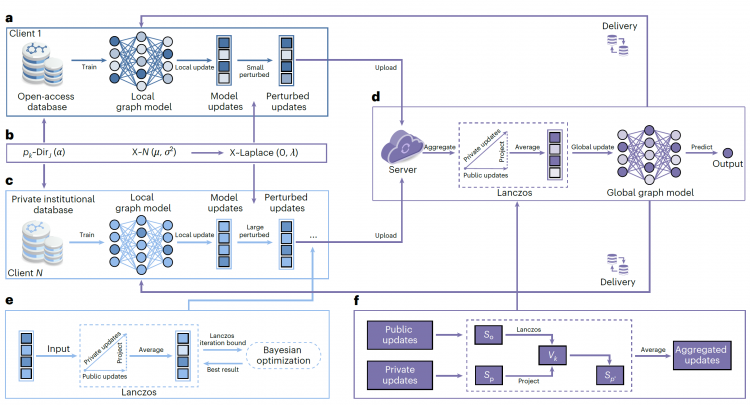
Figure 1. FedLG Method Workflow. The FedLG method applies different intensities of Laplacian perturbation to open data and private data respectively, and utilizes the Lanczos algorithm for projection and weighted average aggregation of global model updates, achieving privacy-preserving training of heterogeneous molecular data under the federated learning framework.
Federated Learning (FL) has emerged in recent years as a promising framework for jointly training global models using data stored across multiple remote clients. Applying FL to molecular discovery means that research institutions can collaboratively perform molecular learning tasks while keeping sensitive data decentralized and private. However, multi-party collaborative model training methods based on traditional FL algorithms may still carry risks of information leakage without strong privacy protection. Attackers may infer critical details of original data or even recover data by analyzing model updates or intermediate transmissions.
II. Research Content: Design and Performance Validation of the FedLG Method
(A) Method Design and Implementation
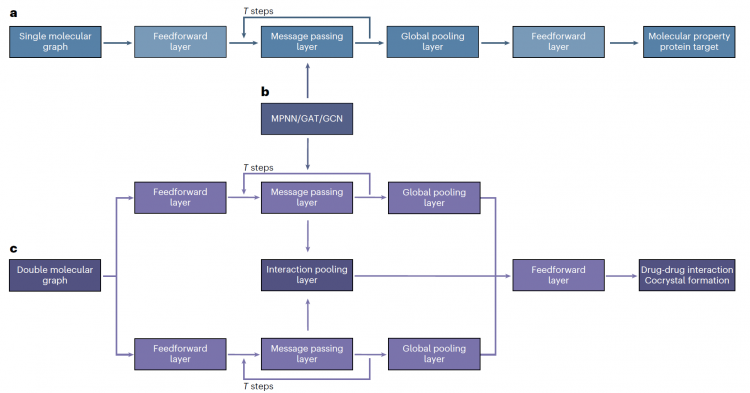
Figure 2. GNN Architecture. Three graph neural network architectures and their message passing mechanisms (MPNN, GAT, GCN) used in the FedLG method for processing single-molecule graph input and dual-molecule graph comparison.
FedLG fully leverages the accessibility, low sensitivity, and public availability of open-access molecular databases, adopting a heterogeneous differential privacy mechanism. It uses the Lanczos algorithm to project updates from high-privacy private institutional databases into the subspace of open-access database model updates. Subsequently, weighted average aggregation is performed to construct the global model, taking into account the different privacy budgets assigned to private institutions and open-access molecular databases. To accommodate various graph models, the research team employed three classical graph neural network designs for single-molecule and dual-molecule graph inputs: MPNN (Message-Passing Neural Network), GAT (Graph Attention Network), and GCN (Graph Convolutional Network).
(B) Effectiveness Validation
The research team compared FedLG with multiple representative federated baseline methods across 18 different datasets. In molecular property prediction and protein target virtual screening tasks, FedLG outperformed all federated baseline methods on eight datasets from MoleculeNet and seven datasets from LIT-PCBA. In DDI and cocrystal material formation prediction tasks, FedLG achieved improvements of 33.9%, 42.3%, and 20.8% on the DrugBank, BIOSNAP, and CoCrystal datasets, respectively. Experimental results showed that FedLG demonstrated strong noise resistance across various noise levels, with only minor fluctuations in model performance as label change noise rates increased from 10% to 20%.
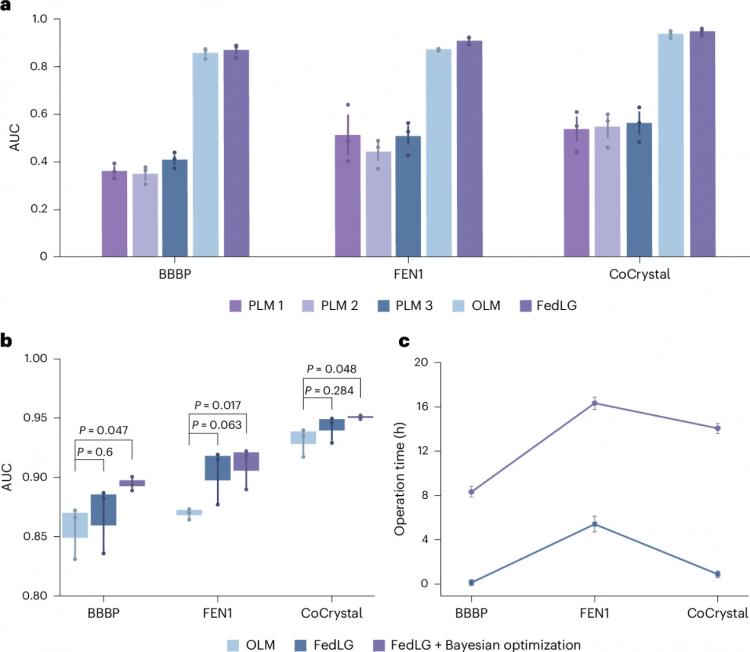
Figure 3. Comparison of FedLG Method Using MPNN Model in Terms of Effectiveness, Performance, and Computational Efficiency
To demonstrate the effectiveness of FedLG, the research team conducted leave-one-client experiments. Results showed that when a single private institutional molecular database was removed, model performance was essentially unaffected; whereas removing the open-access molecular database led to a significant decrease in model performance, highlighting the critical importance of open-access databases. The research team also incorporated Bayesian optimization into FedLG to adaptively determine the optimal number of Lanczos iterations, further improving global model performance.
With the rapid development of AI and large language models, data privacy and security issues have become increasingly prominent. FedLG uses the Lanczos algorithm to capture key information from local model updates of open-access molecular databases, then projects local model updates from private institutional databases onto the extracted features, ensuring that the global model receives better aggregated updates. Extensive experimental results demonstrate that FedLG can perform various molecular discovery tasks under strict privacy protection conditions while maintaining robust behavior and effectively aggregating heterogeneous data. FedLG demonstrates its application potential in privacy-preserving scenarios involving cross-organizational and cross-institutional collaboration, with its ability to collaborate across institutions without compromising sensitive molecular information, promising to transform the research paradigm of molecular discovery.
Paper Link:https://www.nature.com/articles/s42256-026-01184-1

Yuen Wu
Professor and PhD Supervisor at the Department of Applied Chemistry, University of Science and Technology of China (USTC), concurrently a part-time professor at the First Affiliated Hospital of USTC, and a recipient of the Chang Jiang Distinguished Professor title from the Ministry of Education. In recent years, Professor Wu has focused on the rational design of single-atom and cluster catalysts and their exploration in industrial applications, achieving remarkable accomplishments in the precise activation of "chemical bonds" in small molecules in the fields of energy and catalysis. In 2015, he received funding from the National Natural Science Foundation of China's Excellent Young Scientists Fund; in 2017, he was funded by the National Key R&D Program of China's Nanotechnology Special Youth Project and served as Principal Investigator; in 2018, he received the Chinese Chemical Society's Nanochemistry Rising Star Award; in 2019, he received the Chinese Chemical Society's Young Chemist Award; in 2020, he received the Fok Ying Tung Young Teacher Award; in 2023, he received the Optics Valley Achievement Transformation Award; and in 2025, he received the Chinese Chemical Society-Royal Society of Chemistry Young Chemist Award.
In recent years, as corresponding author, he has published more than 170 academic papers in mainstream international journals, including Nat. Mach. Intell.(1), Nat. Nanotechnol.(1), Nat. Catal.(3), Nat. Synth.(1), PNAS(2), J. Am. Chem. Soc.(16), Angew. Chem. Int. Ed.(20), Nat. Commun.(14), Adv. Mater.(10), Joule(2), etc. His papers have been cited more than 33,000 times, with an h-index of 85. He was selected as a Clarivate Highly Cited Researcher from 2020 to 2024. He currently serves as an Editorial Board Member for the journal Industrial Chemistry & Materials, an Editorial Board Member for Science China Materials, a Guest Editor for Small Methods (Single-Atom Catalysis Special Issue), a Youth Editorial Board Member for the Chinese Journal of Inorganic Chemistry, a Youth Editorial Board Member for Chemical Research in Chinese Universities, a Committee Member of the Fuel Cell Branch of the Internal Combustion Engine Association, a Committee Member of the CO2 Division of the Chinese Chemical Society, and a Committee Member of the Nanozyme Division of the Chinese Chemical Society.
中科大团队Nature Machine Intelligence: FedLG
联邦图学习方法实现分子发现的多方协作
优化分子资源利用以加速分子发现进程,需要研究机构和组织之间的协同努力。然而,鉴于每个机构产生的成功和失败分子都具有很高的研究价值,这些发现通常在正式发表或商业化之前被高度保密。这种保密要求对大多数现有方法在严格隐私约束下协同处理异构分布的分子数据提出了巨大挑战。中国科学技术大学团队提出FedLG(联邦学习Lanczos图),一种联邦图学习方法,利用Lanczos算法促进多方协同模型训练,在严格隐私保护条件下实现可靠的预测性能。
一、研究背景:分子数据隐私与协同需求的矛盾
分子发现被认为是推动材料设计、药物发现等相关领域发展的关键组成部分。随着人工智能和实验技术的发展,研究人员现在能够接触到更多独特的分子结构。这些分子通常分散在不同的研究机构或组织中,呈现出异构的数据分布,同时受到严格的隐私和保密约束。开放获取的分子数据库仅代表可用分子知识的一小部分,因为它们无法访问私人研究机构内的高价值和机密分子数据。
图1. FedLG方法流程图。FedLG方法通过对开放数据和隐私数据分别施加不同强度的拉普拉斯扰动,并利用兰索斯算法投影与加权平均聚合全局模型更新,实现了在联邦学习框架下对异构分子数据的隐私保护训练
联邦学习(FL)近年来作为一种有前景的框架出现,用于使用存储在多个远程客户端的数据联合训练全局模型。将FL应用于分子发现意味着研究机构可以在保持敏感数据去中心化和私密性的同时协同执行分子学习任务。然而,基于传统FL算法的多方协同模型训练方法在没有强大隐私保护的情况下仍可能携带信息泄露的风险。攻击者可能通过分析模型更新或中间传输推断原始数据的关键细节,甚至恢复数据。
二、研究内容:FedLG方法的设计与性能验证
(一)方法设计与实现
图2. GNN架构。FedLG方法中用于处理单分子图输入、双分子图对比的三种图神经网络架构及其消息传递机制(MPNN (message-passing neural network)、GAT (graph attention network)、GCN (graph convolutional network))
FedLG充分利用开放获取分子数据库的可访问性、低敏感性和公开性,采用异构差分隐私(heterogeneous differential privacy)机制,使用Lanczos算法将高隐私保护的私人机构数据库的更新投影到开放获取数据库模型更新的子空间中。随后执行加权平均聚合以构建全局模型,同时考虑到分配给私人机构和开放获取分子数据库的不同隐私预算。为适应各种图模型,研究团队使用了MPNN (message-passing neural network)、GAT (graph attention network)、GCN (graph convolutional network)三种经典的图神经网络设计用于单分子和双分子图输入。
(二)有效性验证
研究团队在18个不同的数据集上将FedLG与多个代表性联邦基线方法进行了比较。在分子性质预测和蛋白质靶点虚拟筛选任务中,FedLG在MoleculeNet的八个数据集和LIT-PCBA的七个数据集上均优于所有联邦基线方法。在DDI和共晶材料形成预测任务中,FedLG在DrugBank、BIOSNAP和CoCrystal数据集上分别提升了33.9%、42.3%和20.8%。实验结果显示,FedLG在多种噪声水平下表现出强大的抗噪声能力,随着标签更改噪声率从10%增加到20%,模型性能仅出现轻微波动。
图3. 使用MPNN模型的FedLG方法在有效性、性能及计算效率方面的比较
为证明FedLG的有效性,研究团队进行了留一客户端实验。结果表明,当移除单个私人机构分子数据库时,模型性能基本不受影响;而移除开放获取分子数据库会导致模型性能大幅下降,凸显了开放获取数据库的关键重要性。研究团队还将贝叶斯优化纳入FedLG,以自适应地确定最佳Lanczos迭代次数,进一步提高了全局模型性能。
随着AI和大语言模型的快速发展,数据隐私和安全问题日益突出。FedLG使用Lanczos算法捕获来自开放获取分子数据库的本地模型更新中的关键信息,然后将来自私人机构数据库的本地模型更新投影到提取的特征上,确保全局模型获得更好的聚合更新。广泛的实验结果表明,FedLG可以在严格隐私保护条件下执行多种分子发现任务,同时保持稳健的行为并有效聚合异构数据。FedLG展示了其在涉及跨组织和跨机构协作的隐私保护场景中的应用潜力,在不损害敏感分子信息的情况下跨机构协作的能力,有望改变分子发现的研究范式。
论文链接:https://www.nature.com/articles/s42256-026-01184-1
吴宇恩,中国科学技术大学应用化学系教授及博士生导师,同时担任中国科学技术大学第一附属医院的兼职教授,并被授予教育部长江特聘教授称号。近年来,吴教授专注于单原子和团簇催化剂的理性设计,以及这些催化剂在工业应用中的探索,特别是在能源和催化领域小分子“化学键”的精准活化方面取得了显著成就。2015年获基金委优秀青年基金资助,2017年获国家重点研发计划纳米专项青年项目资助并任首席,2018年获得中国化学会纳米化学新锐奖,2019年获得中国化学会青年化学奖,2020年获得霍英东青年教师奖,2023年获得光谷成果转化奖,2025年获得中国化学会−英国皇家化学会青年化学奖。近年来,以通讯作者在国际主流期刊发表学术论文170余篇,包含 Nat. Mach. Intell.(1), Nat. Nanotechnol.(1), Nat.Catal.(3), Nat. Synth. (1), PNAS (2), J. Am. Chem. Soc.(16), Angew.Chem. Int. Ed. (20), Nat. Commun. (14), Adv. Mater. (10) Joule (2)等,论文总引用33000余次,h-index 85, 2020-2024年入选科睿唯安高被引科学家。目前担任期刊Industrial Chemistry & Materials编委,Science China Materials编委,Small Methods客座编辑(单原子催化专刊),无机化学学报青年编委,Chemical Research in Chinese Universities青年编委,内燃机协会燃料电池分会委员以及中国化学会CO2分会委员,中国化学会纳米酶分会委员。


